Much wider availability of next-generation sequencing (NGS) panels has made it possible to individualize treatment options for a subset of prostate cancer patients with specific tumor markers or alterations. These developments have created a number of new questions, many of which may be answered though the accumulation of clinical and genomic data in a “real-world” dataset.
Sequencing of Therapies
With the evolution of multiple novel treatment options in mCRPC, including novel hormonal agents and PARP inhibitors, as well as the emergence of novel therapeutic classes such as PSMA-targeted radioligand therapies, there is increasingly a wealth of options to offer patients. The PROMISE clinical-genomic platform can help to generate initial data on potential positive and negative predictive biomarkers to guide these types of therapeutic questions.
Natural History & Treatment Options for Novel Molecular Subtypes
As treatment for prostate cancer is increasingly individualized and new biological subsets are identified through the increased use of NGS, it is also important to better define the natural history of these patient populations. The molecular characterization of exceptional responders and non-responders to standard of care therapies will help better define molecular predictive biomarkers. The improved understanding of molecularly defined subsets through hypothesis-generating retrospective studies can help improve the design of biomarker-driven clinical trials of prostate cancer.
Impact of Race, Ethnicity & Socioeconomic Status on Treatments & Outcomes
Many studies focusing on characterizing the molecular alterations in prostate cancer have included a very limited number of ethnically diverse subsets of patients. Understanding how race and ethnicity impacts genomic profile and response to therapy is critical to bridging the disparities gap in men with prostate cancer. Real-world data from this platform can fill this gap.
The inclusion of underrepresented minorities in this dataset is one of the important missions of this consortium. Data from retrospective series can more extensively capture clinical outcomes and other information of more diverse populations, including minority populations underrepresented in clinical trials. The selection for this consortium of diverse clinical sites representing different geographic regions of the United States that serve unique underrepresented patient populations is an important step toward bridging the cancer genomics disparities gap in prostate cancer.
Patient Data
Patient data is captured in a secure RedCap database, with over 1,000 records having been entered as of early 2021. Our goal is to have over 2,500 patient records entered by the end of 2021.
Inclusion Criteria
Inclusion criteria for patients in this dataset consist of having advanced prostate cancer with either mHSPC or castrate-resistant prostate cancer (CRPC). In addition, patients should have undergone prior germline and/or somatic tumor molecular profiling studies including, but not limited to, one or more of the following blood or tissue-based assays:
- tumor DNA sequencing (e.g., cell-free circulating tumor DNA or tumor tissue sequencing)
- germline DNA sequencing and transcript profiling studies (e.g., RNA-seq, qRT-PCR)
In total, consecutive patients with available clinical and genomic data at each site will be included to limit selection bias. Clinical and treatment data is collected from the time of initial prostate cancer diagnosis. All data are completely deidentified without any protected health information. Genomic information entered into the database from both solid tissue and liquid biopsy testing and inclusive of both somatic and germline alterations are formatted to standardized criteria and reviewed by the Bio-Informatics Committee.
All data gathered will be retrospective in nature, through review of medical charts/electronic records, with no direct patient contact.
Examples of Clinical & Genomic Data Collected
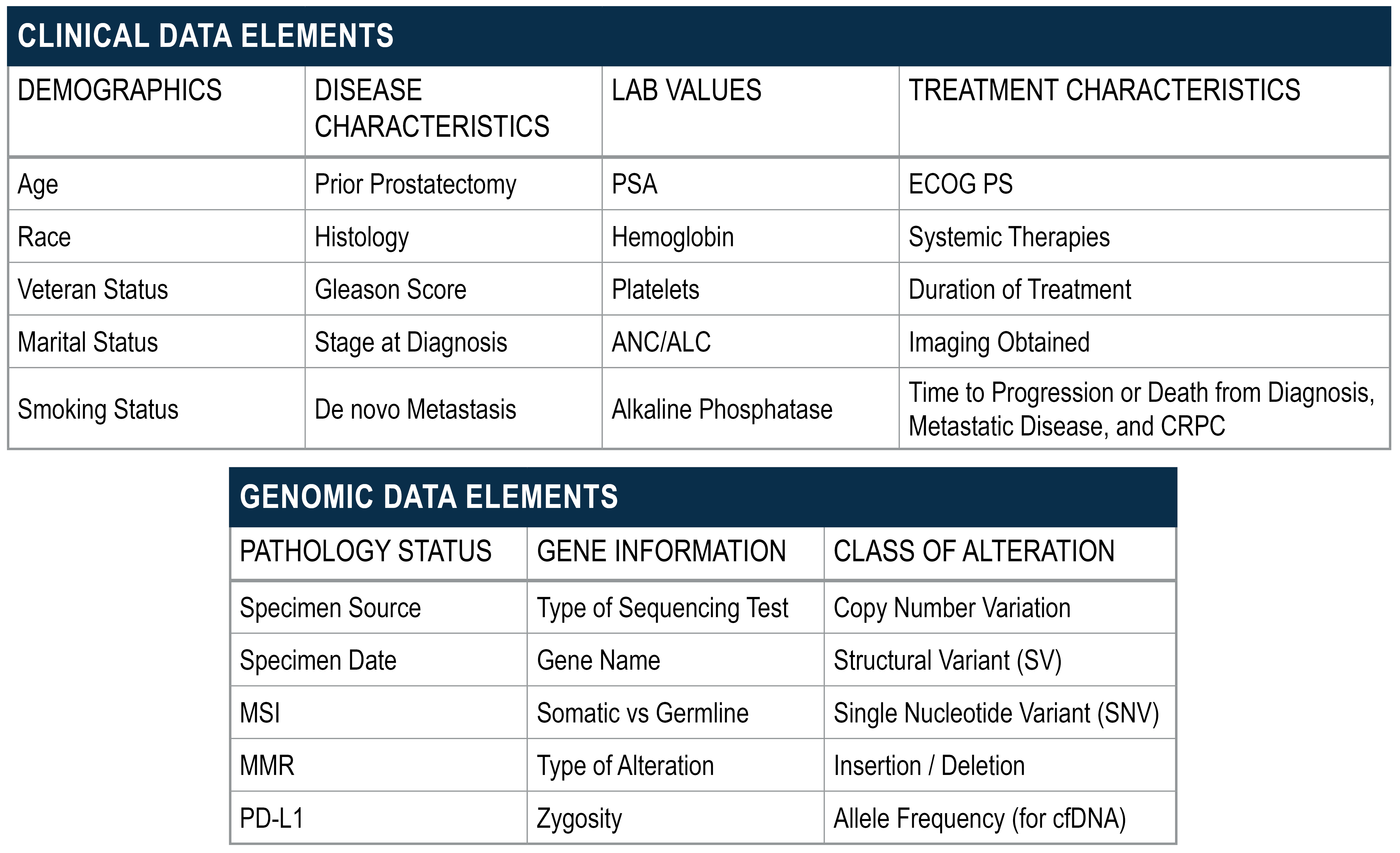
Abstracts & Publications
- Clinical Implications of Wnt Signaling Alterations in Patients (pts) with Advanced Prostate Cancer (aPC) - 2023 ASCO Genitourinary Cancers Symposium Poster Session, February 2023
- Biomarker-Directed Therapy in Black and White Men with Metastatic Castration-Resistant Prostate Cancer (mCRPC) - 2022 ASCO Annual Meeting, June 2022
- Implications of Androgen Receptor (AR) Alterations Identified by Genomic Testing of Tissue and Blood from Advanced Prostate Cancer (aPC) Patients (pts) - 2022 ASCO Genitourinary Cancers Symposium Poster Session, February 2022
- DNA Damaging Therapies in Patients (pts) with Prostate Cancer (PC) and Pathogenic Alterations in Homologous Recombination Repair (HRR) Genes - 2022 ASCO Genitourinary Cancers Symposium Poster Session, February 2022
- PROMISE: A Real-World Clinical-Genomic Database to Address Knowledge Gaps in Prostate Cancer, Prostate Cancer and Prostatic Diseases, August 2021
Grant
- Novel Genomic Prognostic Model in Enzalutamide Treated Prostate Cancer - National Comprehensive Cancer Network (NCCN), Pfizer Global Medical Grants (Pfizer), and Astellas Pharma Global Development



