Showing 1-15 of 66 results

Health Lab
Researchers have used advanced computer algorithms to uncover distinct molecular subgroups of kidney diseases, independent of clinical classifications. These findings have significant implications for personalized treatment approaches.
Health Lab
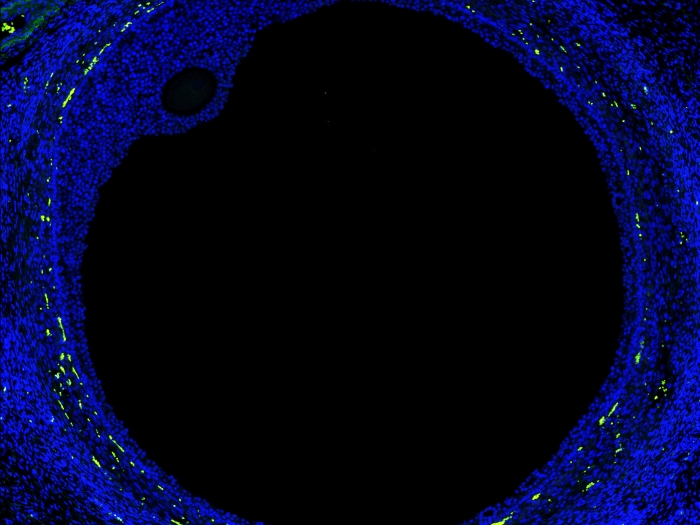
New map of the ovary provides a deeper understanding of how oocytes interact with the surrounding cells during the normal maturation process, and how the function of the follicles may break down in aging or fertility related diseases.
Department News
Stephen C.J. Parker, Ph.D., is the recipient of the 2024 Outstanding Scientific Achievement Award from the American Diabetes Association (ADA).
Department News
Jun Li was inducted into the 2024 Class of the AIMBE College of Fellows.

Department News
The Rajapakse lab receives a new grant.

Health Lab
At-home test can detect tumor DNA fragments in urine samples, providing a non-invasive alternative to traditional blood-based biomarker tests
Department News
Former DCMB PhD student Dr. Shuze Wang was published in Developmental Cell.
Department News
Meet Dr. April Kriebel, a recent graduate from the Welch lab.

Health Lab
An updated rat reference provides more accuracy for research; could help researchers using rat models for the study of DNA, RNA, evolution, or genes linked to disease risks

Health Lab
New study measured continuous heart rate, step count, sleep data and daily mood scores. The researchers also estimated circadian time and time awake from minute-by-minute wearable heart rate and motion measurements.
Department News
Ahmed Elhossiny and Kevin Yang, both Bioinformatics PhD students, have been awarded a Rackham Predoctoral Fellowship.
Department News
DCMB renewed its support of i2b2 tranSMART.
Department News
On March 12, 2024, Elisa Warner defended her dissertation titled "Advancing Clinical Outcome Prediction through Innovative Multimodal and Domain-Generalized AI that Accommodates Limited Data."
Department News
The new Epigenomic Metabolic Medicine center (EM2C) will contribute to understanding how genetic variations contribute to common, complex diseases such as diabetes